Difference between Restriction Endonuclease and Exonuclease
The difference between exonuclease and endonuclease class 12 is one of the most important topics in molecular biology. Both are nucleases, also called molecular scissors, but they act at different sites. The key difference between restriction endonucleases and exonucleases lies in their site of action. Restriction endonucleases (restriction enzymes) cut DNA or RNA internally at specific recognition sites. Exonucleases enzyme remove nucleotides sequentially from the ends of DNA or RNA strands. Understanding the difference between exonuclease and endonuclease class 12 helps students score well in exams and strengthens biotechnology concepts.
This Story also Contains
- What are Restriction Endonuclease and Restriction Exonuclease?
- Difference between Restriction Endonuclease and Exonuclease
- Restriction Endonuclease Enzyme and Its Functions
- Restriction Exonuclease Enzyme and Its Role in DNA Repair
- Types of Restriction Endonuclease (Type I, II, III, and IV)
- Application of Restriction Endonuclease in Genetic Engineering
- Exonuclease Enzyme Types: 5’→3’ and 3’→5’
- Application of Restriction Exonuclease in Replication and Proofreading
- NEET MCQs on Difference Between Exonuclease and Endonuclease (With Answers & Explanations)
- Restriction Enzymes Recommended Video:
The main function of these molecular scissors is to divide the DNA at specific points. These restriction enzymes have major applications in genetic engineering, cloning, DNA replication and repair. This article explains the difference between restriction endonucleases and exonucleases class 12, including definitions, mechanisms, and types of endonucleases with applications. By understanding the functions of restriction enzymes and exonucleases enzymes, students can strengthen their biotechnology principles and processes concepts with NEET-focused MCQs.
What are Restriction Endonuclease and Restriction Exonuclease?
Restriction endonucleases(restriction enzymes) cut the DNA molecules internally at a specific recognition site. Exonucleases remove nucleotides from the ends of DNA or RNA molecules and are involved in genetic engineering and molecular biology. Nucleases are therefore important in the manipulation of nucleic acids because they hydrolyse the phosphodiester bonds within DNA and RNA.
Due to the many roles of nucleases in genetic engineering and molecular biology, it is important to learn about restriction endonucleases, exonucleases, and other nucleases. Among them, restriction endonucleases, which cut DNA at specific sites, are necessary for cloning, gene engineering, and recombinant DNA technology. In contrast, exonucleases are primarily involved in DNA repair as well as replication, ensuring genome stability.
The enzymes often act as molecular scissors as they can divide, splice, or join nucleic acids, which has thoroughly changed molecular biology. It encouraged the creation of techniques like CRISPR and gene therapies shows how these enzymes are valuable in research and industry.
Difference between Restriction Endonuclease and Exonuclease
There are certain differences between the endonuclease and the exonuclease, which separate them from one another. Restriction endonucleases and exonucleases differ in their site of action, mechanism, and applications in molecular biology. Let us differentiate between restriction endonucleases and exonucleases based on the features in the table below:
Features | Restriction Endonucleases | Exonucleases Enzyme |
|---|---|---|
Definition | Enzymes that cut DNA at specific recognition sites | Enzymes that remove nucleotides from the ends of DNA |
Site of Action | Internal DNA sequences | DNA ends (5’ or 3’) |
Mechanism | Recognise and cleave specific DNA sequences | Trim nucleotides sequentially from DNA ends. |
Product obtained | The product obtained after cleavage is an oligonucleotide chain. | The product obtained after exonuclease activity is a monomer of nucleotides. |
Types | Type I, II, III, and IV | 5' to 3' exonucleases, 3' to 5' exonucleases |
Applications | Cloning vector, gene editing, and recombinant DNA | DNA repair, removal of RNA primers, and DNA replication |
Lag Period | It shows a lag period during their activity. | It does not show any lag period. |
Example | EcoRI, HindIII | Exonuclease I, Exonuclease III |
Recognition Sequence | Specific palindromic sequences | Not sequence-specific |
Cutting Pattern | Specific cuts produce sticky or blunt ends | Sequential removal of nucleotides |
Usage in Laboratories | Creating recombinant DNA, restriction mapping | DNA repair studies, degradation of RNA primers |
Restriction Endonuclease Enzyme and Its Functions
Restriction endonuclease, also called restriction enzymes, are the molecular tools that hydrolyse DNA at specific restriction sites that exist within a DNA molecule. These enzymes act as precise scissors and are widely used in genetic engineering because they cut the DNA at specific places for insertion or alteration of genes.
Restriction endonucleases were identified for the first time in the early 1960s during research on the strategies used by bacteria to fight against viral infection. Hamilton Smith and his co-workers named the first restriction enzyme discovered as HindII, and it existed in the bacterium Haemophilus influenzae. This finding transformed molecular biology and was the foundational innovation for constructing genetically tailored modern technology.
Mechanism of action
Recognition sites: Restriction endonucleases come into contact with the target DNAs by the direct complementary base interactions, usually palindromic nucleotide sequences. Once bound, the enzyme causes the breakdown of phosphodiester bonds within the DNA structure, thus causing DNA cleavage and dividing it into two sections.
Cleavage Patterns: Depending on the enzyme, DNA may be cut straight across both strands (producing blunt ends) or at staggered positions (producing sticky ends). For example, EcoRI recognises the sequence 5’ GAATTC 3’ and generates sticky ends, making it a powerful tool in recombinant DNA technology.
Restriction Exonuclease Enzyme and Its Role in DNA Repair
Exonucleases are enzymes that remove nucleotides through endonucleases provided in the nucleic acids of both DNA and RNA. They have important functions related to DNA repair, replication, and RNA processing because they generate and recognise basic sites or any nucleotide lesion.
The enzyme mechanism involves exonucleases participating in the breakdown of a phosphodiester bond of the nucleotide. This results in the release of nucleotides at the terminal of the DNA or RNA strand. They also have polarity depending on which direction of the nucleic acid strand they originate from.
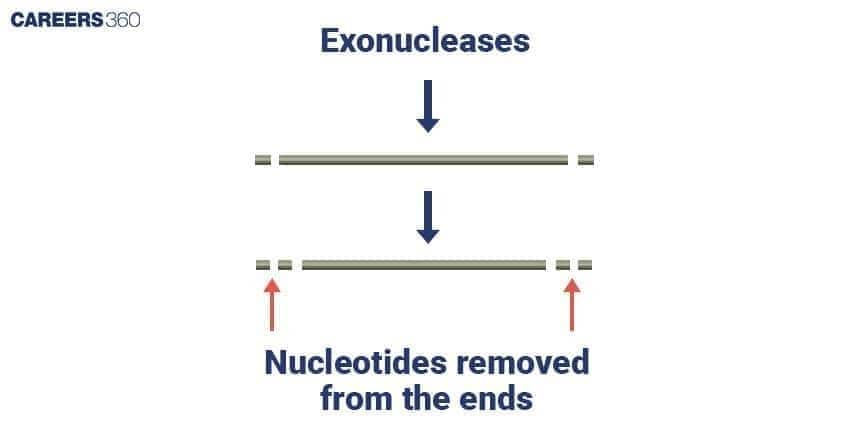
Restriction Exonuclease Diagram
Types of Restriction Endonuclease (Type I, II, III, and IV)
There are four types of restriction endonucleases based on their mechanisms of action and site of cleavage. Each type has distinct properties as given below:
Type I: These enzymes only bind to some DNA sequences, but cut at any position that is arbitrary relative to the binding site.
Type II: These enzymes recognise specific DNA sequences and cut at defined positions within or near the recognition site. Most widely used in molecular biology and biotechnology. Examples: EcoRI, HindIII.
Type III: These enzymes can bind with the DNA sequence, and the DNA gets cut at a fixed distance from the binding site. These require ATP for cleavage.
Type IV: These enzymes cut DNA at particular sites, but for this procedure, they do not need ATP hydrolysis.
Example of Restriction Endonuclease
One of the more commonly known restriction endonucleases is EcoRI. It was the very first restriction enzyme isolated. This cuts DNA at a specific recognition sequence - 5’ GAATTC 3’ that allows it to be an important tool for molecular biology in dealing with DNA manipulation.
Application of Restriction Endonuclease in Genetic Engineering
There are a few really important applications of restriction endonucleases. From genetic engineering and cloning to DNA mapping, it works as a saviour. The application of Restriction endonuclease enzymes is given below:
Genetic engineering: Restriction endonucleases are used to divide the DNA at definite sites so that the foreign DNA fragments can be inserted or the unwanted sequences can be deleted.
Cloning: They are also involved in the formation of recombinant DNA molecules through their ability to make V-shaped cuts at the restriction recognition site of the vector and insert DNA.
DNA mapping: In molecular biology, methods like restriction fragment length polymorphism analysis (RFLP) and Southern blotting, restriction endonuclease enzymes are used to determine the sequence of DNA strands and variations, respectively.
Exonuclease Enzyme Types: 5’→3’ and 3’→5’
Exonucleases are enzymes that remove nucleotides from the ends of DNA or RNA strands. They work in two main directions, depending on the type of exonuclease. There are two main types of exonucleases:
5’ to 3’ exonucleases: These enzymes remove nucleotides one at a time from the 5’ end of the DNA or RNA strand. They plays role in DNA replication and RNA primer removal.
3’ to 5’ exonucleases: These enzymes cut nucleotides off the 3’ end of the DNA or RNA strand in a stepwise manner. These are important for DNA repair and proofreading.
Example of Restriction Exonuclease
One of the common exonuclease enzymes is ExoI. This enzyme degrades single-stranded DNA from the 3' end and is implicated in DNA repair and recombination processes.
Application of Restriction Exonuclease in Replication and Proofreading
Exonucleases are essential enzymes in molecular biology and genetic engineering. They remove nucleotides from the ends of DNA or RNA strands and play a critical role in maintaining genetic material. Exonucleases have multiple applications in molecular biology:
DNA repair: Exonucleases participate in the base excision repair processes of DNA and nucleotide excision repair. They remove the damaged, mismatched nucleotides in the DNA strands.
Degradation of RNA primers: During the replication of DNA, RNA primers are utilised and eventually erased by exonucleases so that DNA polymerase can extend the newly created strand with DNA nucleotide.
Removal of mismatched nucleotides: Exonucleases interact with DNA mismatch repair pathways to help remove nucleotides that fail to pair with the template DNA strand in the right way, thereby maintaining genome stability.
NEET MCQs on Difference Between Exonuclease and Endonuclease (With Answers & Explanations)
Question 1: Restriction endonucleases are enzymes which
Make cuts at specific positions within the DNA molecule
Recognise a specific nucleotide sequence for the binding of DNA ligase
Restrict the action of the enzyme DNA Polymerase
remove nucleotides from the ends of the DNA molecule
Correct Answer: 1) Make cuts at specific positions within the DNA molecule
Explanation: As we learnt in Restriction Endonuclease, these enzymes cleave DNA only within or near the particular base sequence. These sequences are called recognition sites. They are of three types: RE-I, RE-II and RE-III. - wherein the 1st Restriction enzyme (Hind II) was used in RDT.
Hence, the correct answer is option (1) Make cuts at specific positions within the DNA molecule.
Question 2: Nuclease who removes nucleotides from the ends of the DNA in one strand of the duplex.
Exonuclease
Endonuclease
Exo-endo nuclease
None of the above
Correct Answer: 1)Exonuclease
Explanation:
Exonucleases are enzymes that specifically remove nucleotides from the ends of DNA strands. They function by cleaving nucleotides one at a time from either the 5′ end or the 3′ end of a single strand within a DNA duplex. This precise activity makes exonucleases important for various cellular processes, including DNA repair, replication, and degradation of unneeded or damaged DNA.
Hence, the correct answer is option 1)Exonuclease.
Question 3: The term 'molecular scissors' refers to
Recombinant DNA
Restriction enzymes
Taq polymerase
Palindromic nucleotide sequences
Correct Answer: 2) Restriction enzymes
Explanation:
Cleaving enzymes, also known as restriction enzymes or molecular scissors, are used to cleave DNA molecules at specific sites. There are three main types: Exonucleases, which remove nucleotides from the ends of DNA; Endonucleases, which cut DNA at internal sites; and Restriction endonucleases, which recognise and cut DNA at specific sequences known as restriction sites. These enzymes are widely used in molecular biology for DNA manipulation, including tasks such as cloning, DNA mapping, and genetic engineering.
Hence, the correct answer is option 2) Restriction enzymes.
Question 4: A specific recognition sequence identified by endonucleases to make cuts at specific positions within the DNA is :
Degenerate primer sequence
Okazaki sequences
Palindromic Nucleotide Sequences
Poly (A) tail sequences
Correct Answer: 3) Palindromic Nucleotide Sequences
Explanation:
A specific part of DNA that endonucleases can identify and cut is termed a restriction site or recognition sequence. These are like molecular scissors with each enzyme searching for a distinct nucleotide pattern, typically 4-8 bases long. An example is EcoRI, which finds the sequence 5’-GAATTC-3’ and cuts just after the first G. Another is HindIII, which locates 5’-AAGCTT-3’ and snips between the two A's. These enzymes are vital in fields like genetic engineering and molecular biology for creating gene copies, mapping DNA, and other useful lab techniques. They help us to fragment DNA into smaller segments with either sticky or blunt ends, which can be rearranged as necessary for different experiments.
Hence, the correct answer is option 3) Palindromic Nucleotide sequences.
Restriction Enzymes Recommended Video:
Frequently Asked Questions (FAQs)
Restriction endonucleases: Cut DNA at specific recognition sites within the DNA strand.
Exonucleases: Remove nucleotides one at a time from the ends of DNA strands.
Nucleases are enzymes that cut nucleic acids. Endonucleases cleave at internal sites within DNA/RNA, while exonucleases remove nucleotides stepwise from the ends.
The exonuclease enzyme removes RNA primers and mismatched nucleotides during replication. This ensures accurate DNA copying and genome stability.
Restriction endonucleases are called molecular scissors because they cut DNA at exact recognition sites, producing sticky or blunt ends useful in cloning and recombinant DNA technology.
The first restriction endonuclease isolated was HindII, which was discovered in 1970. It cuts DNA at specific recognition sites, making it a pivotal discovery in the field of molecular biology.

